Portfolio

My classwork from bimm143
Class 18: Pertusssis Mini Project
Sam Fisher
Background
Pertussis (aka whooping cough) is a common lung infection caused by the bacteria B. Pertussis.
This can infect people of any age, but it is most lethal to infants due to their smaller airways. https://www.cdc.gov/pertussis/php/surveillance/pertussis-cases-by-year.html?CDC_AAref_Val=https://www.cdc.gov/pertussis/surv-reporting/cases-by-year.html The CDC tracks the number of reported cases in the US: We can “scrape” this data with the datapasta package
cdc <- data.frame(
year = c(1922L,1923L,1924L,1925L,
1926L,1927L,1928L,1929L,1930L,1931L,
1932L,1933L,1934L,1935L,1936L,
1937L,1938L,1939L,1940L,1941L,1942L,
1943L,1944L,1945L,1946L,1947L,
1948L,1949L,1950L,1951L,1952L,
1953L,1954L,1955L,1956L,1957L,1958L,
1959L,1960L,1961L,1962L,1963L,
1964L,1965L,1966L,1967L,1968L,1969L,
1970L,1971L,1972L,1973L,1974L,
1975L,1976L,1977L,1978L,1979L,1980L,
1981L,1982L,1983L,1984L,1985L,
1986L,1987L,1988L,1989L,1990L,
1991L,1992L,1993L,1994L,1995L,1996L,
1997L,1998L,1999L,2000L,2001L,
2002L,2003L,2004L,2005L,2006L,2007L,
2008L,2009L,2010L,2011L,2012L,
2013L,2014L,2015L,2016L,2017L,2018L,
2019L,2020L,2021L,2022L,2023L, 2024L, 2025L),
cases = c(107473,164191,165418,152003,
202210,181411,161799,197371,
166914,172559,215343,179135,265269,
180518,147237,214652,227319,103188,
183866,222202,191383,191890,109873,
133792,109860,156517,74715,69479,
120718,68687,45030,37129,60886,
62786,31732,28295,32148,40005,
14809,11468,17749,17135,13005,6799,
7717,9718,4810,3285,4249,3036,
3287,1759,2402,1738,1010,2177,2063,
1623,1730,1248,1895,2463,2276,
3589,4195,2823,3450,4157,4570,
2719,4083,6586,4617,5137,7796,6564,
7405,7298,7867,7580,9771,11647,
25827,25616,15632,10454,13278,
16858,27550,18719,48277,28639,32971,
20762,17972,18975,15609,18617,
6124,2116,3044,7063, 22538, 21996)
)
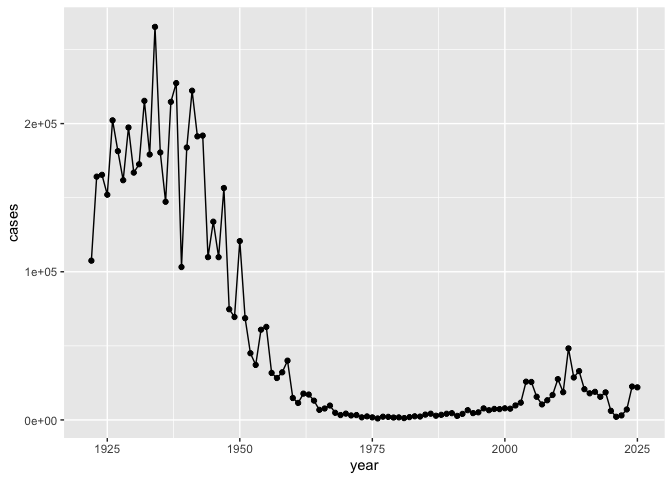
Q. Make a plot of
yearvscases
library(ggplot2)
ggplot(cdc, aes(x = year, y = cases)) +
geom_point() + geom_line()
Q. Add some major milestones including the first wP vaccine roll-out (1946), the switch to the newer aP vaccine (1996), the COVID virus (2020)
ggplot(cdc, aes(x = year, y = cases)) +
geom_point() +
geom_line() +
geom_vline(xintercept = 1946, linetype = "dashed", color = "blue") +
geom_vline(xintercept = 1996, linetype = "dashed", color = "red") +
geom_vline(xintercept = 2020, linetype = "dashed", color = "green")
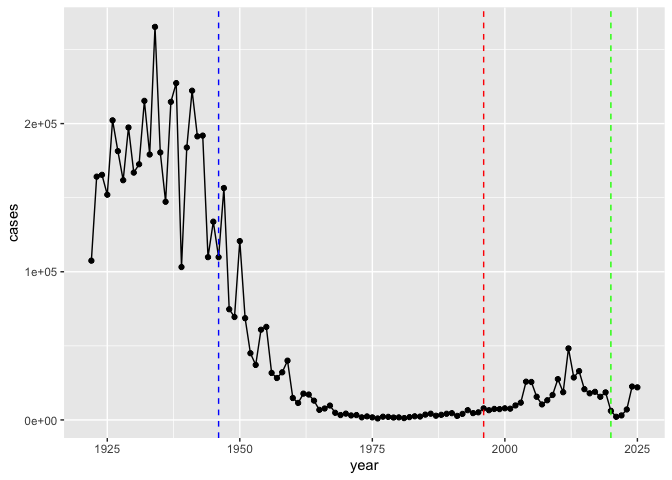
There were high case numbers in the pre 1940s followed by a sharp decline after the introduction of the whole-cell pertussis vaccine in
- The spike in the mid 2000s is likely associated with anti-vaccine movements in the media that have caused people to believe vaccines are bad for their health.
There were high case numbers pre 1940s followed by a sharp decline after the introduction of the whole-cell pertussis vaccine in 1946. Case numbers remained fairly low and constant until the mid 2000s when the ap vaccine was introduced. Countries such as Japan who switched two years before the U.S. show a similar pattern but two years earlier. Other countries that switched later also show the same pattern but later. It appears that the ap vaccine wears off faster.
Why is this vaccine-preventable disease on the upswing? To answer this question we need to investigate the mechanisms underlying waning protection against pertussis. This requires evaluation of pertussis-specific immune responses over time in wP and aP vaccinated individuals.
CMI-PB Data
Computational Models of Immunity - Pertussis Boost project aims to provide the scientific community with this very information.
They make their data available vis JSON formati returning API. We can
read this in R with the read_json() function from the jsonlite
package:
library(jsonlite)
subject <- read_json("https://www.cmi-pb.org/api/v5_1/subject", simplifyVector = TRUE)
head(subject)
subject_id infancy_vac biological_sex ethnicity race
1 1 wP Female Not Hispanic or Latino White
2 2 wP Female Not Hispanic or Latino White
3 3 wP Female Unknown White
4 4 wP Male Not Hispanic or Latino Asian
5 5 wP Male Not Hispanic or Latino Asian
6 6 wP Female Not Hispanic or Latino White
year_of_birth date_of_boost dataset
1 1986-01-01 2016-09-12 2020_dataset
2 1968-01-01 2019-01-28 2020_dataset
3 1983-01-01 2016-10-10 2020_dataset
4 1988-01-01 2016-08-29 2020_dataset
5 1991-01-01 2016-08-29 2020_dataset
6 1988-01-01 2016-10-10 2020_dataset
Q. How mant wP and aP individuals are in this table?
table(subject$infancy_vac)
aP wP
87 85
Q. What is the biological sex breakdown
table(subject$biological_sex)
Female Male
112 60
Q. In terms of race and gender is this dataset representative of the US population
table(subject$race, subject$biological_sex)
Female Male
American Indian/Alaska Native 0 1
Asian 32 12
Black or African American 2 3
More Than One Race 15 4
Native Hawaiian or Other Pacific Islander 1 1
Unknown or Not Reported 14 7
White 48 32
Let’s read some more database tables:
specimen <- read_json("http://cmi-pb.org/api/v5_1/specimen", simplifyVector = TRUE)
ab_titer <- read_json("http://cmi-pb.org/api/v5_1/plasma_ab_titer", simplifyVector = TRUE)
head(specimen)
specimen_id subject_id actual_day_relative_to_boost
1 1 1 -3
2 2 1 1
3 3 1 3
4 4 1 7
5 5 1 11
6 6 1 32
planned_day_relative_to_boost specimen_type visit
1 0 Blood 1
2 1 Blood 2
3 3 Blood 3
4 7 Blood 4
5 14 Blood 5
6 30 Blood 6
head(ab_titer)
specimen_id isotype is_antigen_specific antigen MFI MFI_normalised
1 1 IgE FALSE Total 1110.21154 2.493425
2 1 IgE FALSE Total 2708.91616 2.493425
3 1 IgG TRUE PT 68.56614 3.736992
4 1 IgG TRUE PRN 332.12718 2.602350
5 1 IgG TRUE FHA 1887.12263 34.050956
6 1 IgE TRUE ACT 0.10000 1.000000
unit lower_limit_of_detection
1 UG/ML 2.096133
2 IU/ML 29.170000
3 IU/ML 0.530000
4 IU/ML 6.205949
5 IU/ML 4.679535
6 IU/ML 2.816431
To analyze this data we need to first “join” (merge/link) the different tables so we ahve all ther data in one place not spread accross different tables.
We can use the *_join() family of functions from dplyr to do this
library(dplyr)
Attaching package: 'dplyr'
The following objects are masked from 'package:stats':
filter, lag
The following objects are masked from 'package:base':
intersect, setdiff, setequal, union
meta <- inner_join(subject, specimen)
Joining with `by = join_by(subject_id)`
head(meta)
subject_id infancy_vac biological_sex ethnicity race
1 1 wP Female Not Hispanic or Latino White
2 1 wP Female Not Hispanic or Latino White
3 1 wP Female Not Hispanic or Latino White
4 1 wP Female Not Hispanic or Latino White
5 1 wP Female Not Hispanic or Latino White
6 1 wP Female Not Hispanic or Latino White
year_of_birth date_of_boost dataset specimen_id
1 1986-01-01 2016-09-12 2020_dataset 1
2 1986-01-01 2016-09-12 2020_dataset 2
3 1986-01-01 2016-09-12 2020_dataset 3
4 1986-01-01 2016-09-12 2020_dataset 4
5 1986-01-01 2016-09-12 2020_dataset 5
6 1986-01-01 2016-09-12 2020_dataset 6
actual_day_relative_to_boost planned_day_relative_to_boost specimen_type
1 -3 0 Blood
2 1 1 Blood
3 3 3 Blood
4 7 7 Blood
5 11 14 Blood
6 32 30 Blood
visit
1 1
2 2
3 3
4 4
5 5
6 6
abdata <- inner_join(ab_titer, meta)
Joining with `by = join_by(specimen_id)`
head(abdata)
specimen_id isotype is_antigen_specific antigen MFI MFI_normalised
1 1 IgE FALSE Total 1110.21154 2.493425
2 1 IgE FALSE Total 2708.91616 2.493425
3 1 IgG TRUE PT 68.56614 3.736992
4 1 IgG TRUE PRN 332.12718 2.602350
5 1 IgG TRUE FHA 1887.12263 34.050956
6 1 IgE TRUE ACT 0.10000 1.000000
unit lower_limit_of_detection subject_id infancy_vac biological_sex
1 UG/ML 2.096133 1 wP Female
2 IU/ML 29.170000 1 wP Female
3 IU/ML 0.530000 1 wP Female
4 IU/ML 6.205949 1 wP Female
5 IU/ML 4.679535 1 wP Female
6 IU/ML 2.816431 1 wP Female
ethnicity race year_of_birth date_of_boost dataset
1 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
2 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
3 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
4 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
5 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
6 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
actual_day_relative_to_boost planned_day_relative_to_boost specimen_type
1 -3 0 Blood
2 -3 0 Blood
3 -3 0 Blood
4 -3 0 Blood
5 -3 0 Blood
6 -3 0 Blood
visit
1 1
2 1
3 1
4 1
5 1
6 1
Q. What antibody isotypes are measured for these patients?
table(abdata$isotype)
IgE IgG IgG1 IgG2 IgG3 IgG4
6698 7265 11993 12000 12000 12000
Q. What antigens are reported?
table(abdata$antigen)
ACT BETV1 DT FELD1 FHA FIM2/3 LOLP1 LOS Measles OVA
1970 1970 6318 1970 6712 6318 1970 1970 1970 6318
PD1 PRN PT PTM Total TT
1970 6712 6712 1970 788 6318
Let’s focus on the IgG isotype and make a plot of MFI_normalized for all antigens.
igg <- abdata |>
filter(isotype == "IgG")
head(igg)
specimen_id isotype is_antigen_specific antigen MFI MFI_normalised
1 1 IgG TRUE PT 68.56614 3.736992
2 1 IgG TRUE PRN 332.12718 2.602350
3 1 IgG TRUE FHA 1887.12263 34.050956
4 19 IgG TRUE PT 20.11607 1.096366
5 19 IgG TRUE PRN 976.67419 7.652635
6 19 IgG TRUE FHA 60.76626 1.096457
unit lower_limit_of_detection subject_id infancy_vac biological_sex
1 IU/ML 0.530000 1 wP Female
2 IU/ML 6.205949 1 wP Female
3 IU/ML 4.679535 1 wP Female
4 IU/ML 0.530000 3 wP Female
5 IU/ML 6.205949 3 wP Female
6 IU/ML 4.679535 3 wP Female
ethnicity race year_of_birth date_of_boost dataset
1 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
2 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
3 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
4 Unknown White 1983-01-01 2016-10-10 2020_dataset
5 Unknown White 1983-01-01 2016-10-10 2020_dataset
6 Unknown White 1983-01-01 2016-10-10 2020_dataset
actual_day_relative_to_boost planned_day_relative_to_boost specimen_type
1 -3 0 Blood
2 -3 0 Blood
3 -3 0 Blood
4 -3 0 Blood
5 -3 0 Blood
6 -3 0 Blood
visit
1 1
2 1
3 1
4 1
5 1
6 1
ggplot(igg) +
aes(MFI_normalised, antigen) + geom_boxplot()
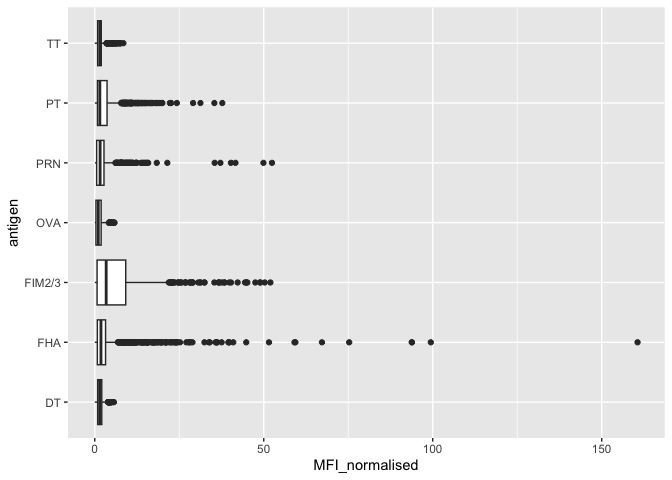
Q. Is there a difference for aP vs wP individuals with these values?
ggplot(igg) +
aes(MFI_normalised, antigen, col=infancy_vac) + geom_boxplot()
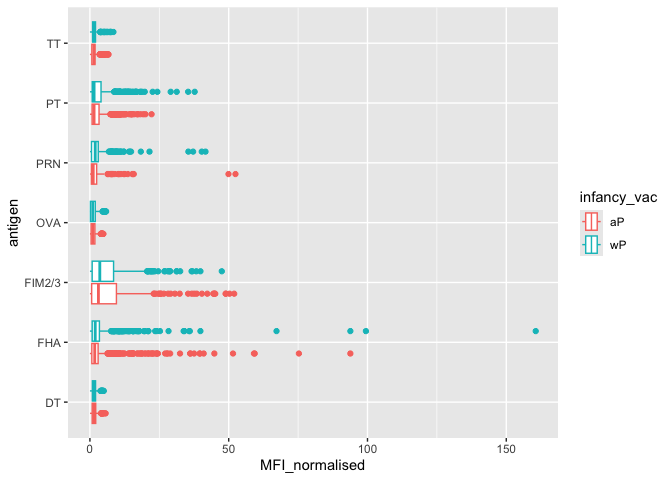
Q. Is there a temrol response - i.e. do values increase or decrease over time?
ggplot(igg) +
aes(MFI_normalised, antigen, col=infancy_vac) + geom_boxplot() + facet_wrap(~visit)
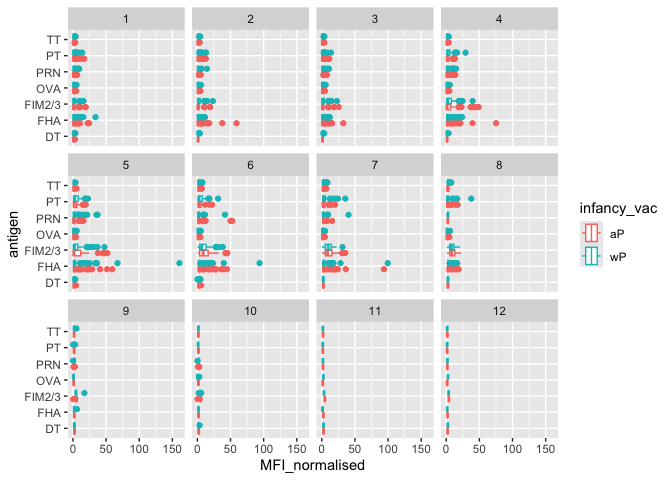
Focus on “PT” Pertusis Toxin antigen
pt.igg.21 <- igg |> filter(antigen == "PT",
dataset == "2021_dataset")
ggplot(pt.igg.21) +
aes(planned_day_relative_to_boost,
MFI_normalised,
col=infancy_vac,
group = subject_id) +
geom_point() + geom_line()
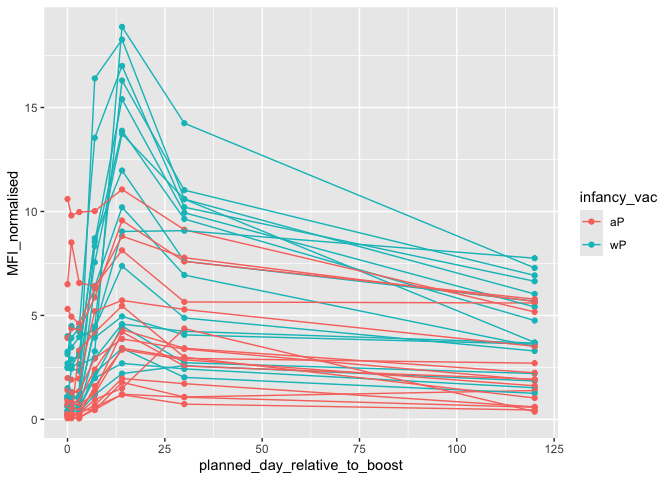
geom_vline(xintercept = 14, lty = "dashed")
mapping: xintercept = ~xintercept
geom_vline: na.rm = FALSE
stat_identity: na.rm = FALSE
position_identity
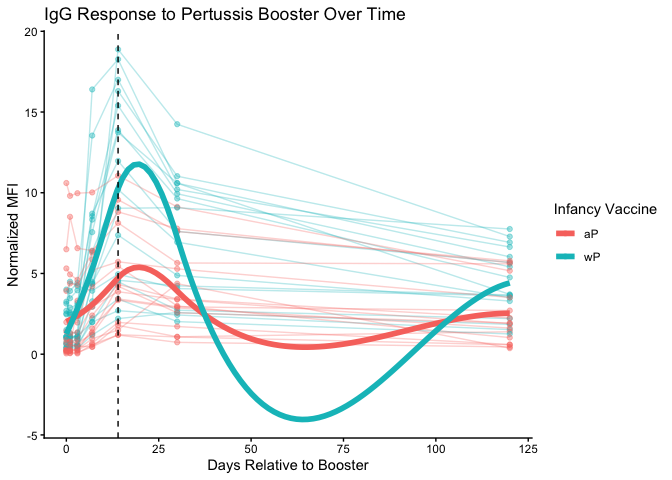
ggplot(pt.igg.21) +
aes(planned_day_relative_to_boost,
MFI_normalised,
col = infancy_vac,
group = subject_id) +
geom_point(alpha = 0.4) +
geom_line(alpha = 0.3) +
geom_smooth(aes(group = infancy_vac), se = FALSE, linewidth = 2) +
geom_vline(xintercept = 14, lty = "dashed") +
labs(
title = "IgG Response to Pertussis Booster Over Time",
x = "Days Relative to Booster",
y = "Normalized MFI",
color = "Infancy Vaccine"
) +
theme_classic()
`geom_smooth()` using method = 'loess' and formula = 'y ~ x'